| NRAS | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
 | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | NRAS, ALPS4, CMNS, N-ras, NCMS, NRAS1, NS6, Neuroblastoma RAS viral oncogene homolog, NRAS proto-oncogene, GTPase | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 164790 MGI: 97376 HomoloGene: 55661 GeneCards: NRAS | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
NRAS is an enzyme that in humans is encoded by the NRAS gene. It was discovered by a small team of researchers led by Robin Weiss at the Institute of Cancer Research in London.[5][6] It was the third RAS gene to be discovered, and was named NRAS, for its initial identification in human neuroblastoma cells.
Function
The N-ras proto-oncogene is a member of the Ras gene family. It is mapped on chromosome 1, and it is activated in HL60, a promyelocytic leukemia line. The order of nearby genes is as follows: cen—CD2—NGFB—NRAS—tel.
The mammalian Ras gene family consists of the Harvey and Kirsten Ras genes (HRAS and KRAS), an inactive pseudogene of each (c-Hras2 and c-Kras1) and the N-Ras gene. They differ significantly only in the C-terminal 40 amino acids. These Ras genes have GTP/GDP binding and GTPase activity, and their normal function may be as G-like regulatory proteins involved in the normal control of cell growth.
The N-Ras gene specifies two main transcripts of 2 kb and 4.3 kb. The difference between the two transcripts is a simple extension through the termination site of the 2 kb transcript. The N-Ras gene consists of seven exons (-I, I, II, III, IV, V, VI). The smaller 2 kb transcript contains the VIa exon, and the larger 4.3 kb transcript contains the VIb exon which is just a longer form of the VIa exon. Both transcripts encode identical proteins as they differ only the 3′ untranslated region.[7]
Mutations
Mutations which change amino acid residues 12, 13 or 61 activate the potential of N-ras to transform cultured cells and are implicated in a variety of human tumors[7] e.g. melanoma.
As a drug target
Binimetinib (MEK162) has had a phase III clinical trial for NRAS Q61 mutant melanoma.[8]
References
- 1 2 3 GRCh38: Ensembl release 89: ENSG00000213281 - Ensembl, May 2017
- 1 2 3 GRCm38: Ensembl release 89: ENSMUSG00000027852 - Ensembl, May 2017
- ↑ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ Marshall CJ, Hall A, Weiss RA (September 1982). "A transforming gene present in human sarcoma cell lines". Nature. 299 (5879): 171–3. Bibcode:1982Natur.299..171M. doi:10.1038/299171a0. PMID 6287287. S2CID 4342747.
- ↑ Shimizu K, Goldfarb M, Perucho M, Wigler M (January 1983). "Isolation and preliminary characterization of the transforming gene of a human neuroblastoma cell line". PNAS. 80 (2): 383–7. Bibcode:1983PNAS...80..383S. doi:10.1073/pnas.80.2.383. PMC 393381. PMID 6300838.
- 1 2 "Entrez Gene: NRAS neuroblastoma RAS viral (v-ras) oncogene homolog".
- ↑ Study Comparing the Efficacy of MEK162 Versus Dacarbazine in Unresectable or Metastatic NRAS Mutation-positive Melanoma
Further reading
- McCormick F (1996). "Ras-related proteins in signal transduction and growth control". Mol. Reprod. Dev. 42 (4): 500–6. doi:10.1002/mrd.1080420419. PMID 8607982. S2CID 6507743.
- van Elsas A, Scheibenbogen C, van der Minne C, et al. (1998). "UV-induced N-ras mutations are T-cell targets in human melanoma". Melanoma Res. 7 (Suppl 2): S107–13. doi:10.1097/00008390-199708001-00017. PMID 9578425. S2CID 19219505.
- Dracopoli NC, Meisler MH (1990). "Mapping the human amylase gene cluster on the proximal short arm of chromosome 1 using a highly informative (CA)n repeat" (PDF). Genomics. 7 (1): 97–102. doi:10.1016/0888-7543(90)90523-W. hdl:2027.42/28584. PMID 1692298.
- Yuasa Y, Kamiyama T, Kato M, et al. (1990). "Transforming genes from familial adenomatous polyposis patient cells detected by a tumorigenicity assay". Oncogene. 5 (4): 589–96. PMID 1970154.
- Hancock JF, Magee AI, Childs JE, Marshall CJ (1989). "All ras proteins are polyisoprenylated but only some are palmitoylated". Cell. 57 (7): 1167–77. doi:10.1016/0092-8674(89)90054-8. PMID 2661017. S2CID 19292550.
- Hall A, Brown R (1985). "Human N-ras: cDNA cloning and gene structure". Nucleic Acids Res. 13 (14): 5255–68. doi:10.1093/nar/13.14.5255. PMC 321863. PMID 2991860.
- Hirai H, Tanaka S, Azuma M, et al. (1986). "Transforming genes in human leukemia cells". Blood. 66 (6): 1371–8. doi:10.1182/blood.V66.6.1371.1371. PMID 2998510.
- Neri A, Knowles DM, Greco A, et al. (1988). "Analysis of RAS oncogene mutations in human lymphoid malignancies". Proc. Natl. Acad. Sci. U.S.A. 85 (23): 9268–72. Bibcode:1988PNAS...85.9268N. doi:10.1073/pnas.85.23.9268. PMC 282720. PMID 3057505.
- Nitta N, Ochiai M, Nagao M, Sugimura T (1987). "Amino-acid substitution at codon 13 of the N-ras oncogene in rectal cancer in a Japanese patient". Jpn. J. Cancer Res. 78 (1): 21–6. PMID 3102434.
- Raybaud F, Noguchi T, Marics I, et al. (1988). "Detection of a low frequency of activated ras genes in human melanomas using a tumorigenicity assay". Cancer Res. 48 (4): 950–3. PMID 3276402.
- Hirai H, Kobayashi Y, Mano H, et al. (1987). "A point mutation at codon 13 of the N-ras oncogene in myelodysplastic syndrome". Nature. 327 (6121): 430–2. Bibcode:1987Natur.327..430H. doi:10.1038/327430a0. PMID 3295562. S2CID 4270769.
- Gambke C, Hall A, Moroni C (1985). "Activation of an N-ras gene in acute myeloblastic leukemia through somatic mutation in the first exon". Proc. Natl. Acad. Sci. U.S.A. 82 (3): 879–82. Bibcode:1985PNAS...82..879G. doi:10.1073/pnas.82.3.879. PMC 397150. PMID 3856237.
- Padua RA, Barrass NC, Currie GA (1985). "Activation of N-ras in a human melanoma cell line". Mol. Cell. Biol. 5 (3): 582–5. doi:10.1128/mcb.5.3.582. PMC 366752. PMID 3887133.
- Brown R, Marshall CJ, Pennie SG, Hall A (1984). "Mechanism of activation of an N-ras gene in the human fibrosarcoma cell line HT1080". EMBO J. 3 (6): 1321–6. doi:10.1002/j.1460-2075.1984.tb01970.x. PMC 557516. PMID 6086315.
- Yuasa Y, Gol RA, Chang A, et al. (1984). "Mechanism of activation of an N-ras oncogene of SW-1271 human lung carcinoma cells". Proc. Natl. Acad. Sci. U.S.A. 81 (12): 3670–4. Bibcode:1984PNAS...81.3670Y. doi:10.1073/pnas.81.12.3670. PMC 345280. PMID 6587382.
- Taparowsky E, Shimizu K, Goldfarb M, Wigler M (1983). "Structure and activation of the human N-ras gene". Cell. 34 (2): 581–6. doi:10.1016/0092-8674(83)90390-2. PMID 6616621. S2CID 39787991.
- Mitchell EL, Jones D, White GR, et al. (1995). "Determination of the gene order of the three loci CD2, NGFB, and NRAS at human chromosome band 1p13 and refinement of their localisation at the subband level by fluorescence in situ hybridisation". Cytogenet. Cell Genet. 70 (3–4): 183–5. doi:10.1159/000134028. PMID 7789166.
- Kodaki T, Woscholski R, Hallberg B, et al. (1995). "The activation of phosphatidylinositol 3-kinase by Ras". Curr. Biol. 4 (9): 798–806. doi:10.1016/S0960-9822(00)00177-9. PMID 7820549. S2CID 20836474.
- Rodriguez-Viciana P, Warne PH, Vanhaesebroeck B, et al. (1996). "Activation of phosphoinositide 3-kinase by interaction with Ras and by point mutation". EMBO J. 15 (10): 2442–51. doi:10.1002/j.1460-2075.1996.tb00602.x. PMC 450176. PMID 8665852.
External links


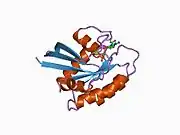


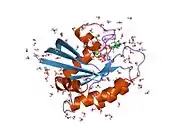
![1iaq: C-H-RAS P21 PROTEIN MUTANT WITH THR 35 REPLACED BY SER (T35S) COMPLEXED WITH GUANOSINE-5'-[B,G-IMIDO] TRIPHOSPHATE](../I/PDB_1iaq_EBI.jpg.webp)
![1jah: H-RAS P21 PROTEIN MUTANT G12P, COMPLEXED WITH GUANOSINE-5'-[BETA,GAMMA-METHYLENE] TRIPHOSPHATE AND MAGNESIUM](../I/PDB_1jah_EBI.jpg.webp)
![1jai: H-RAS P21 PROTEIN MUTANT G12P, COMPLEXED WITH GUANOSINE-5'-[BETA,GAMMA-METHYLENE] TRIPHOSPHATE AND MANGANESE](../I/PDB_1jai_EBI.jpg.webp)